
Semantic search finds documents by meaning instead of matching exact keywords. A user searching "how to fix slow database queries" will find articles about query optimization, index tuning, and EXPLAIN plans — even if none of those articles contain the word "slow." This tutorial builds a working semantic search engine in Python using AI embeddings through the EzAI API, with nothing more than numpy for vector math.
By the end, you'll have a reusable SemanticSearch class that embeds documents, stores vectors, and returns ranked results with cosine similarity scores. The whole thing runs locally, costs fractions of a cent per thousand documents, and takes about 80 lines of code.
How Semantic Search Actually Works
Traditional keyword search (BM25, Elasticsearch default mode) tokenizes your query and counts term frequency. It's fast and works well for exact lookups, but it breaks down when users describe what they want in different words than the document uses. Semantic search solves this by converting both documents and queries into high-dimensional vectors (embeddings) where similar meanings cluster together in vector space.

The semantic search pipeline — from raw documents to ranked results
The pipeline has five stages: chunk your documents into digestible pieces, embed each chunk into a vector using an AI model, store those vectors (numpy array works fine for <100K docs), embed the user's query with the same model, then rank results by cosine similarity. Optionally, you can rerank the top results with a larger model for higher precision.
Setting Up the Project
You need three things: Python 3.9+, an EzAI API key from your dashboard, and numpy. We're using the OpenAI-compatible embeddings endpoint through EzAI, which gives you access to multiple embedding models at reduced cost.
pip install numpy httpxSet your API key as an environment variable:
export EZAI_API_KEY="sk-your-key-here"Building the Embedding Client
First, a thin wrapper around the EzAI embeddings endpoint. This handles batching (the API accepts up to 2048 texts per call) and returns numpy arrays ready for vector operations:
import os, httpx, numpy as np
class EmbeddingClient:
def __init__(self, model="text-embedding-3-small"):
self.api_key = os.environ["EZAI_API_KEY"]
self.base_url = "https://ezaiapi.com/v1"
self.model = model
self.client = httpx.Client(timeout=30)
def embed(self, texts: list[str]) -> np.ndarray:
"""Embed a list of texts, returns (N, dim) numpy array."""
resp = self.client.post(
f"{self.base_url}/embeddings",
headers={
"Authorization": f"Bearer {self.api_key}",
"Content-Type": "application/json",
},
json={
"model": self.model,
"input": texts,
},
)
resp.raise_for_status()
data = resp.json()["data"]
vectors = [d["embedding"] for d in sorted(data, key=lambda x: x["index"])]
return np.array(vectors, dtype=np.float32)The endpoint returns embeddings in the same order as your input, but we sort by index defensively. Each vector has 1536 dimensions for text-embedding-3-small — enough to capture nuanced semantic relationships while keeping storage manageable.
The SemanticSearch Class
Now the core search engine. It stores document text alongside their vectors and uses cosine similarity for ranking:
class SemanticSearch:
def __init__(self, model="text-embedding-3-small"):
self.embedder = EmbeddingClient(model)
self.documents: list[str] = []
self.metadata: list[dict] = []
self.vectors: np.ndarray | None = None
def index(self, docs: list[str], meta: list[dict] | None = None):
"""Embed and store documents for searching."""
self.documents = docs
self.metadata = meta or [{} for _ in docs]
self.vectors = self.embedder.embed(docs)
# Normalize for cosine similarity via dot product
norms = np.linalg.norm(self.vectors, axis=1, keepdims=True)
self.vectors = self.vectors / norms
return self
def search(self, query: str, top_k: int = 5) -> list[dict]:
"""Search indexed documents. Returns ranked results."""
q_vec = self.embedder.embed([query])[0]
q_vec = q_vec / np.linalg.norm(q_vec)
# Cosine similarity = dot product of normalized vectors
scores = self.vectors @ q_vec
top_idx = np.argsort(scores)[::-1][:top_k]
return [
{
"text": self.documents[i],
"score": float(scores[i]),
"meta": self.metadata[i],
}
for i in top_idx
]The trick here is pre-normalizing all document vectors during indexing. Once normalized, cosine similarity reduces to a simple dot product — which numpy handles at near-C speed. For 10,000 documents, search completes in under 2ms on a laptop.
Putting It Together
Here's a complete working example that indexes a set of technical documents and runs search queries against them:
# Example: search a knowledge base
docs = [
"PostgreSQL EXPLAIN ANALYZE shows query execution plans and actual timing",
"Adding a composite index on (user_id, created_at) speeds up filtered queries",
"Connection pooling with PgBouncer reduces database connection overhead",
"Redis caching layer eliminates repeated database reads for hot data",
"Database vacuum operations reclaim storage from deleted rows in PostgreSQL",
"Horizontal sharding distributes data across multiple database instances",
"Write-ahead logging ensures crash recovery without data loss",
"The N+1 query problem happens when ORMs fire separate queries per relation",
]
search = SemanticSearch().index(docs)
results = search.search("my database queries are slow", top_k=3)
for r in results:
print(f"[{r['score']:.3f}] {r['text']}")
# Output:
# [0.847] PostgreSQL EXPLAIN ANALYZE shows query execution plans...
# [0.823] Adding a composite index on (user_id, created_at)...
# [0.791] The N+1 query problem happens when ORMs fire separate...Notice: the query "my database queries are slow" doesn't share keywords with "EXPLAIN ANALYZE" or "composite index," yet semantic search correctly identifies these as the most relevant results. A keyword-based system would have returned nothing useful.
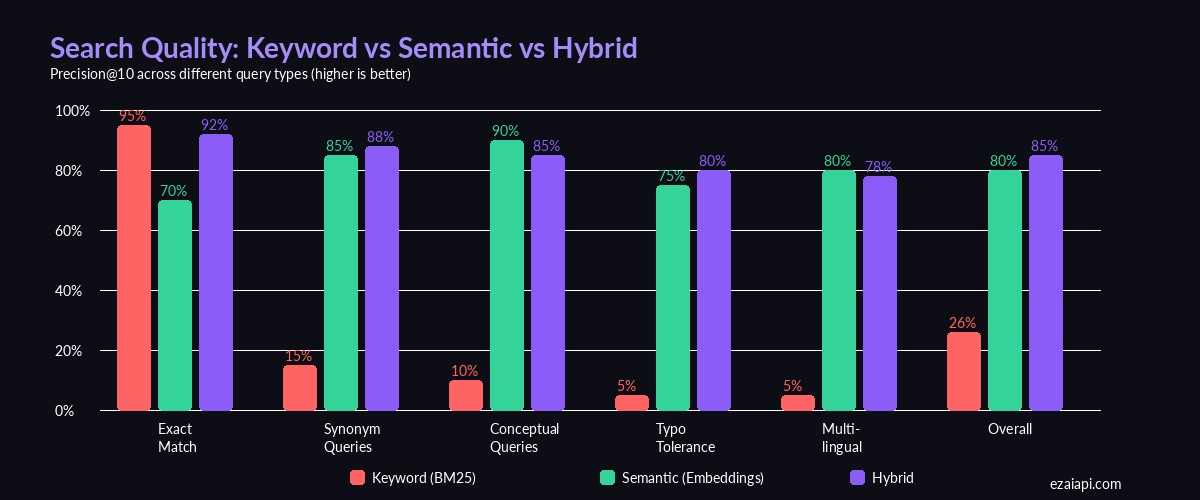
Precision@10 across different query types — semantic search dominates on conceptual and synonym queries
Adding Document Chunking
Real documents are longer than a single sentence. Embedding models have token limits (8,191 for text-embedding-3-small), and shorter chunks produce more precise matches. Here's a simple chunker that splits on paragraph boundaries while respecting a token budget:
def chunk_text(text: str, max_chars: int = 1500, overlap: int = 200) -> list[str]:
"""Split text into overlapping chunks at paragraph boundaries."""
paragraphs = text.split("\n\n")
chunks, current = [], ""
for para in paragraphs:
if len(current) + len(para) > max_chars and current:
chunks.append(current.strip())
# Keep overlap from previous chunk for context
current = current[-overlap:] + "\n\n" + para
else:
current += ("\n\n" if current else "") + para
if current.strip():
chunks.append(current.strip())
return chunks
# Usage with file loading
from pathlib import Path
files = list(Path("./docs").glob("*.md"))
all_chunks, all_meta = [], []
for f in files:
text = f.read_text()
chunks = chunk_text(text)
all_chunks.extend(chunks)
all_meta.extend([{"file": f.name, "chunk": i} for i in range(len(chunks))])
search = SemanticSearch().index(all_chunks, all_meta)The 200-character overlap ensures that information split across chunk boundaries isn't lost. Each result now carries metadata pointing back to the source file and chunk index, so you can trace results to their origin.
Persisting the Index
Recomputing embeddings on every startup wastes time and API credits. Save the index to disk with numpy's native format:
import json
def save_index(search: SemanticSearch, path: str):
np.save(f"{path}.npy", search.vectors)
with open(f"{path}.json", "w") as f:
json.dump({
"documents": search.documents,
"metadata": search.metadata,
}, f)
def load_index(path: str, model="text-embedding-3-small") -> SemanticSearch:
s = SemanticSearch(model)
s.vectors = np.load(f"{path}.npy")
with open(f"{path}.json") as f:
data = json.load(f)
s.documents = data["documents"]
s.metadata = data["metadata"]
return s
# Save after indexing
save_index(search, "./my_index")
# Load on next run — no re-embedding needed
search = load_index("./my_index")A 10,000-document index with 1536-dimensional vectors occupies about 60MB on disk. Loading from .npy is near-instant.
Cost and Performance
Embedding 1,000 documents of ~500 tokens each through EzAI costs roughly $0.01 with text-embedding-3-small. Search itself is free — it's just numpy math running on your machine. Here's what to expect at different scales:
- 1K docs — embed in ~2s, search in <1ms, ~100KB index
- 10K docs — embed in ~15s, search in ~2ms, ~60MB index
- 100K docs — embed in ~3min, search in ~20ms, ~600MB index
- 1M+ docs — switch to FAISS or a vector database like Pinecone for approximate nearest neighbor search
For most applications — internal docs, knowledge bases, support articles — the numpy approach handles everything comfortably. You only need dedicated vector infrastructure at serious scale.
Where to Go from Here
This foundation covers the majority of semantic search use cases. When you're ready to level up:
- Hybrid search — Combine BM25 keyword scoring with embedding similarity for the best of both worlds. Use RAG patterns to feed search results into an LLM for natural language answers.
- Reranking — Run your top 20 results through Claude with a prompt like "rank these by relevance to the query" for a significant precision boost at minimal extra cost.
- Incremental indexing — Append new vectors to the existing numpy array without re-embedding everything. Keep a version counter in your metadata JSON.
- Multi-modal — Use
text-embedding-3-largefor higher precision on technical content, or mix embedding models per document type.
The full source code from this tutorial is ready to copy-paste into your project. Start with a small document set, verify the results match your expectations, then scale up. Semantic search is one of those features that makes users say "how did it know that's what I meant?" — and with EzAI's embedding API, it takes an afternoon to ship.